PDF) Genome-Wide Association Study Demonstrates the Role Played by the CD226 Gene in Rasa Aragonesa Sheep Reproductive Seasonality
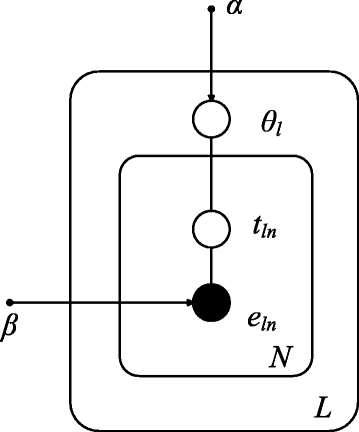
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text
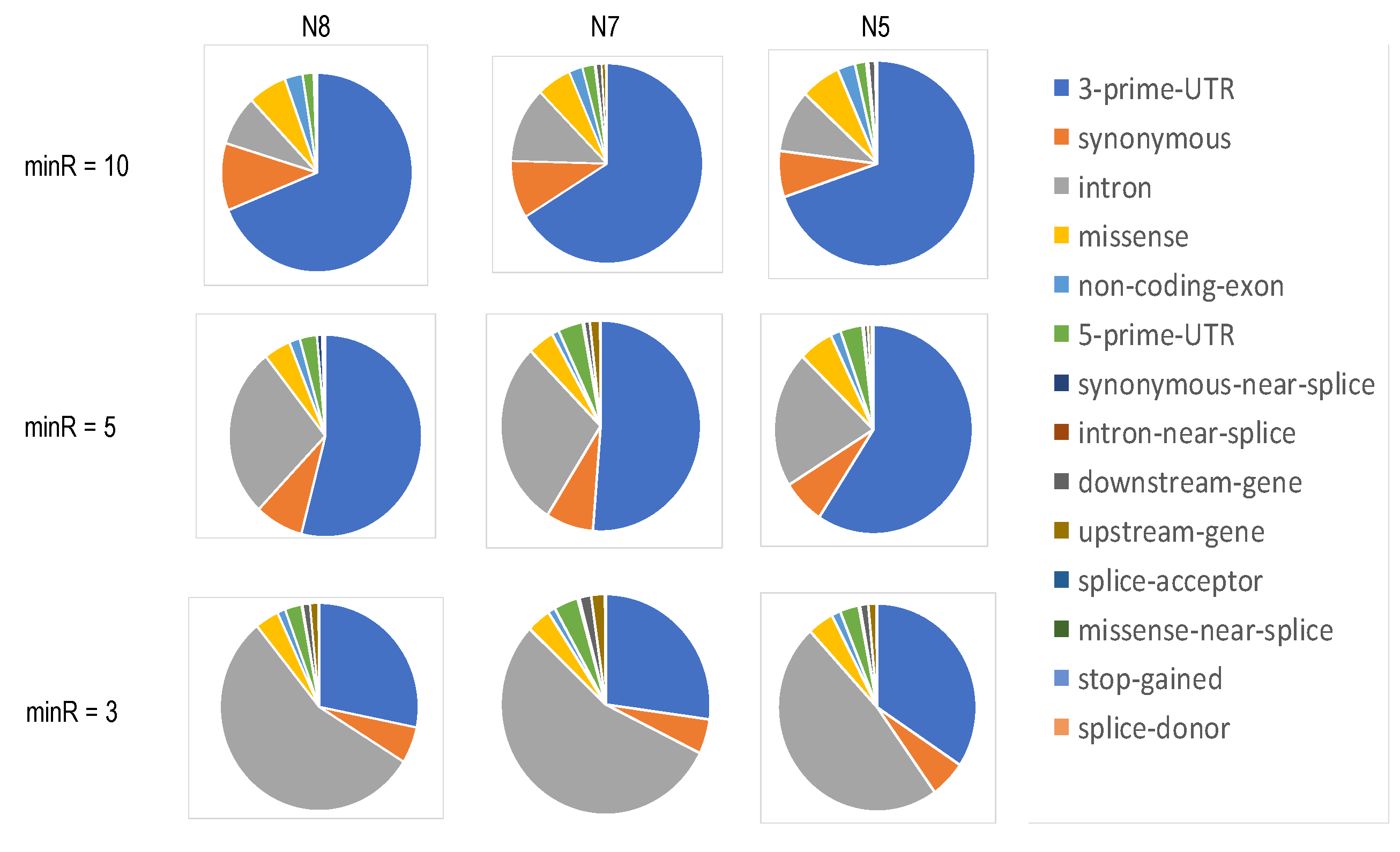
Genes | Free Full-Text | Estimating the Allele-Specific Expression of SNVs From 10× Genomics Single-Cell RNA-Sequencing Data | HTML
ChiTaH: a fast and accurate tool for identifying known human chimeric sequences from high-throughput sequencing data

Mapping of candidate region for chordoma development to 1p36.13 by LOH analysis - Riva - 2003 - International Journal of Cancer - Wiley Online Library
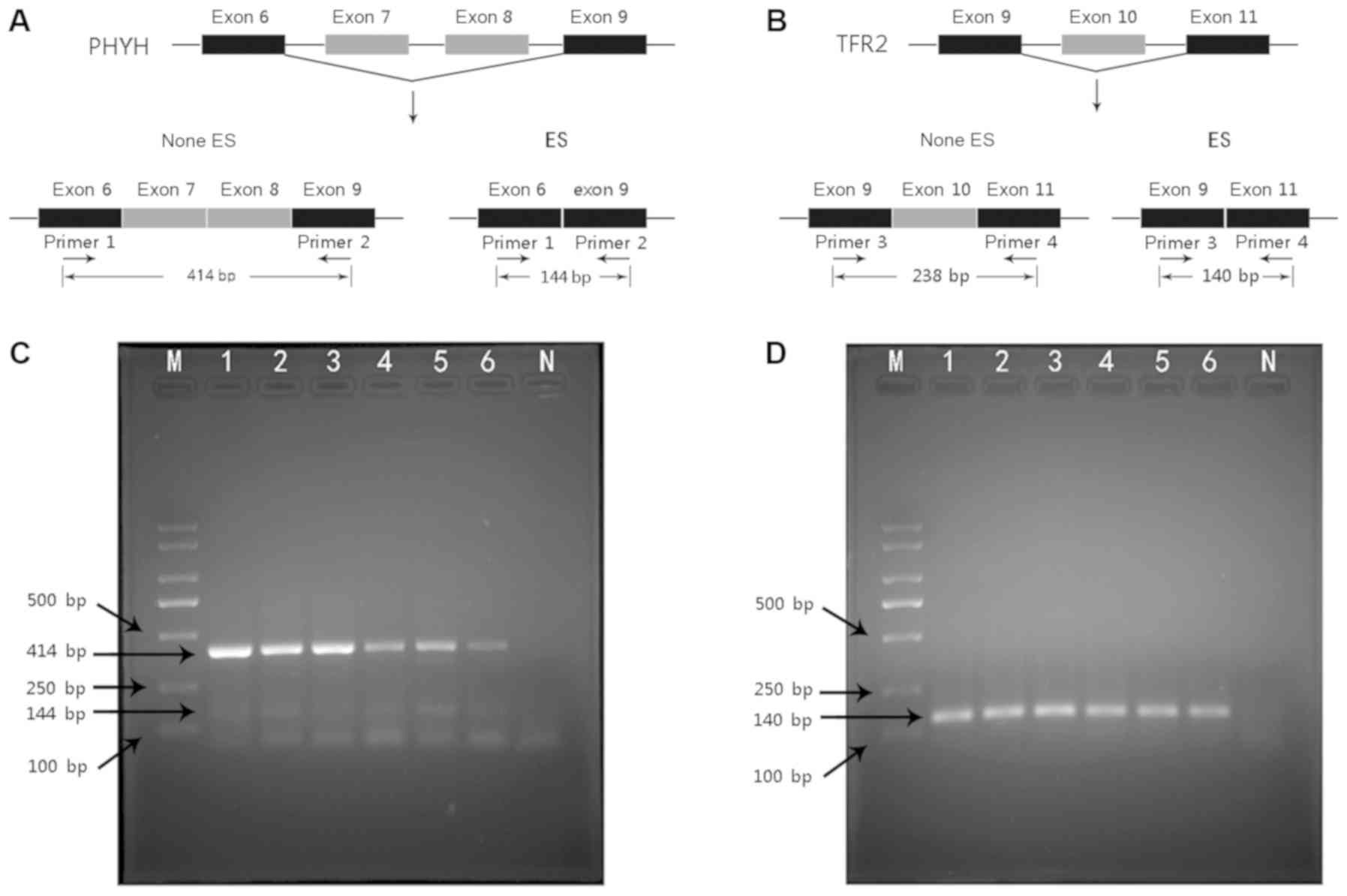
Construction of a model to predict the prognosis of patients with cholangiocarcinoma using alternative splicing events

PDF) The exon quantification pipeline (EQP): A comprehensive approach to the quantification of gene, exon and junction expression from RNA-seq data

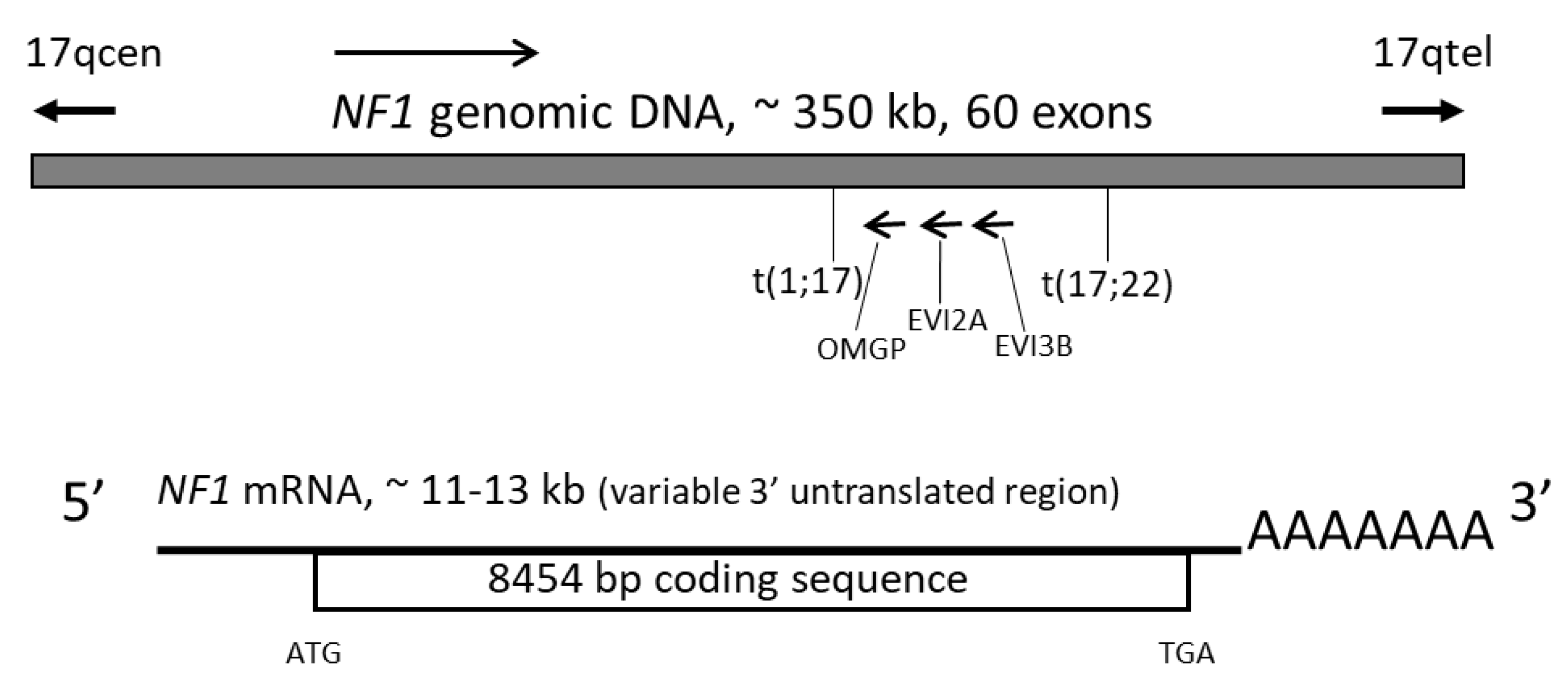
Optimized Lentiviral Vectors for HIV Gene Therapy: Multiplexed Expression of Small RNAs and Inclusion of MGMTP140K Drug Resistance Gene - ScienceDirect
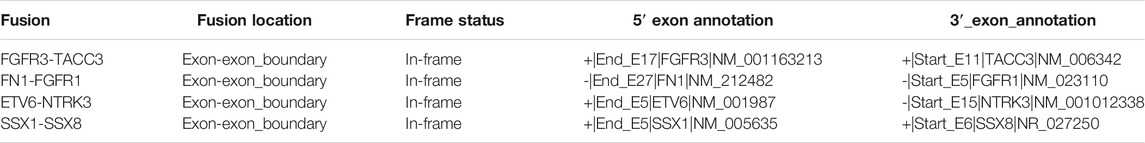
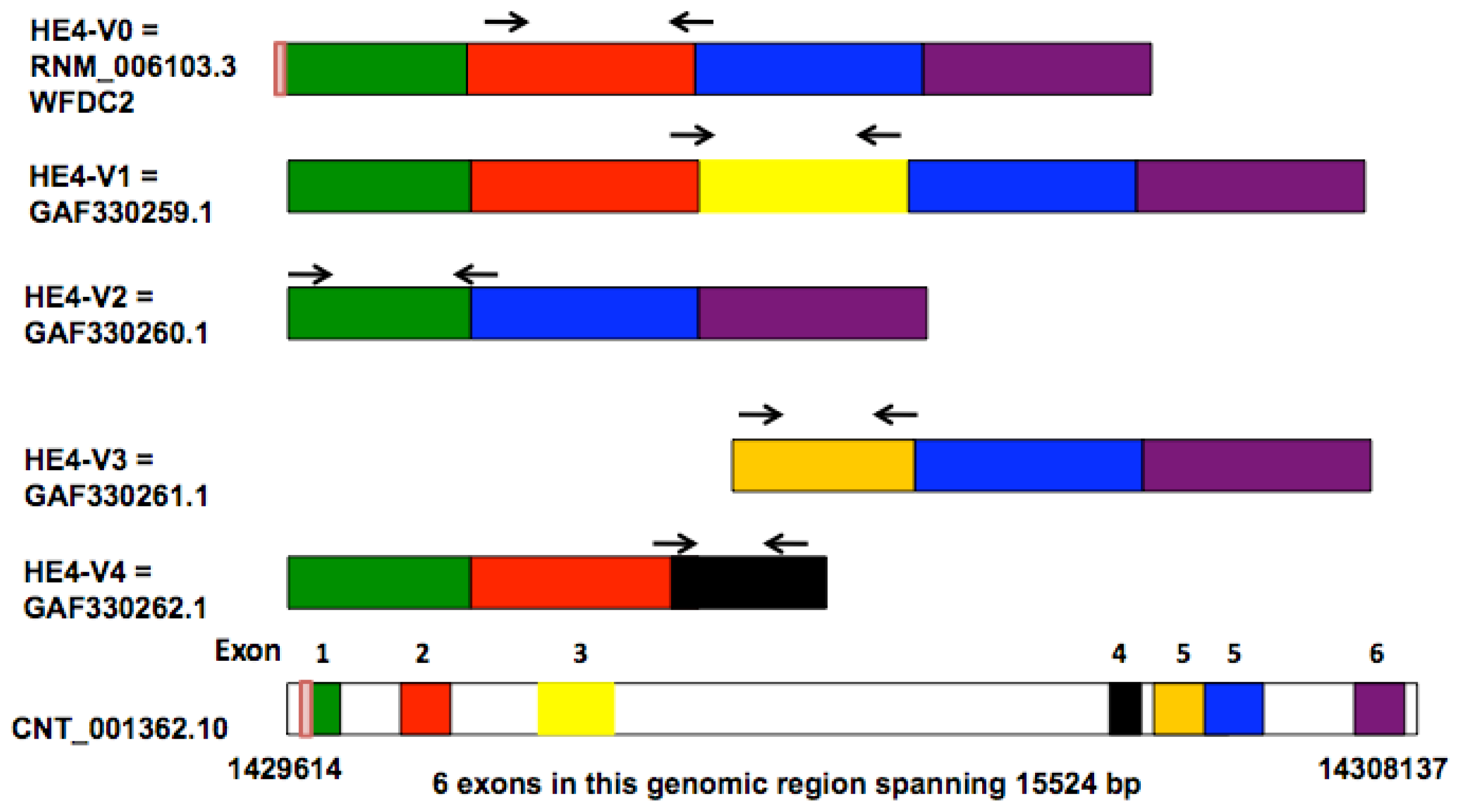
Frontiers | SeekFusion - A Clinically Validated Fusion Transcript Detection Pipeline for PCR-Based Next-Generation Sequencing of RNA
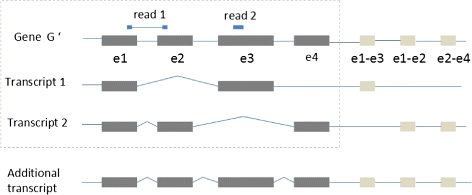
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text