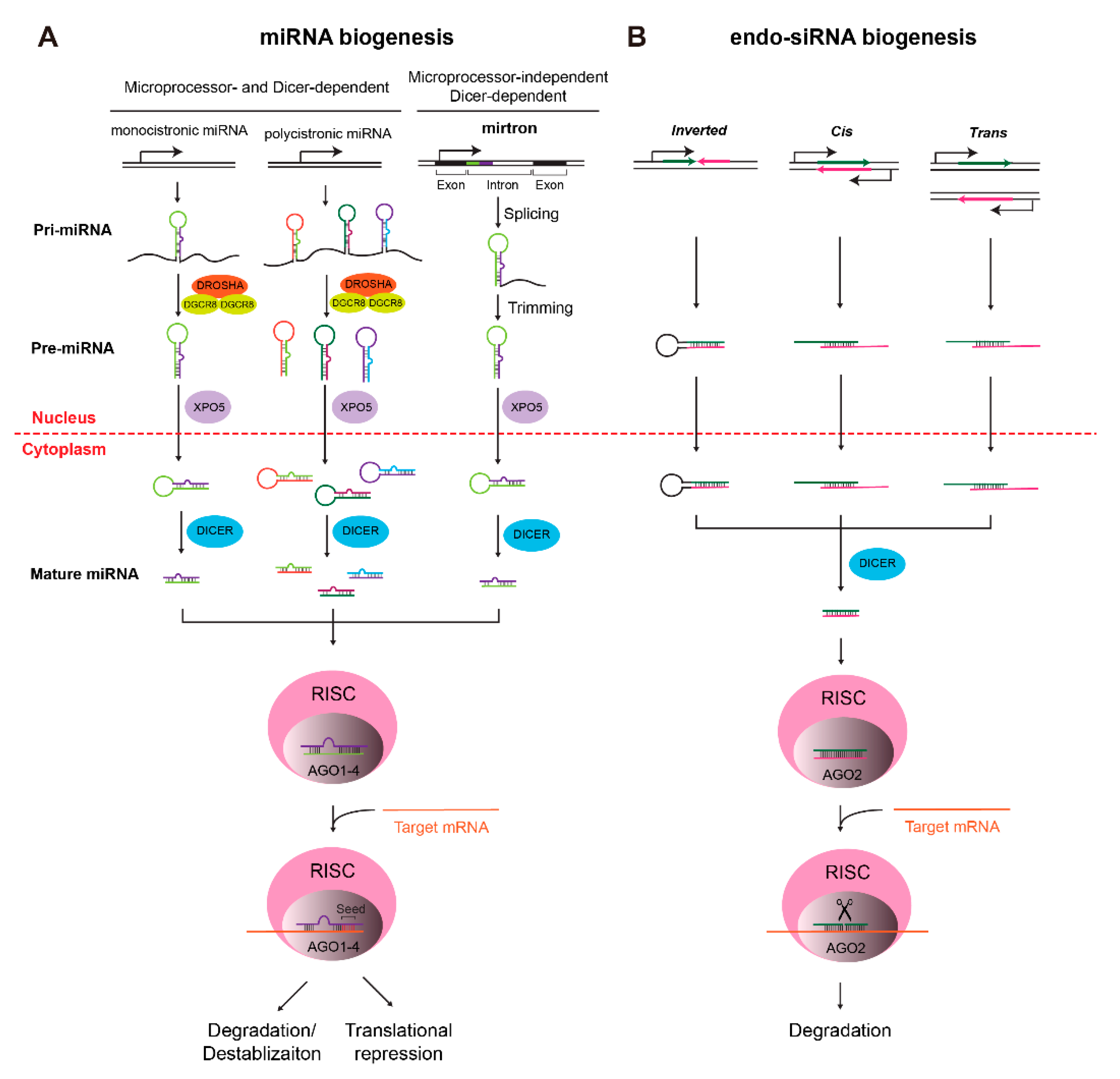
IJMS | Free Full-Text | Roles of MicroRNAs in Establishing and Modulating Stem Cell Potential | HTML
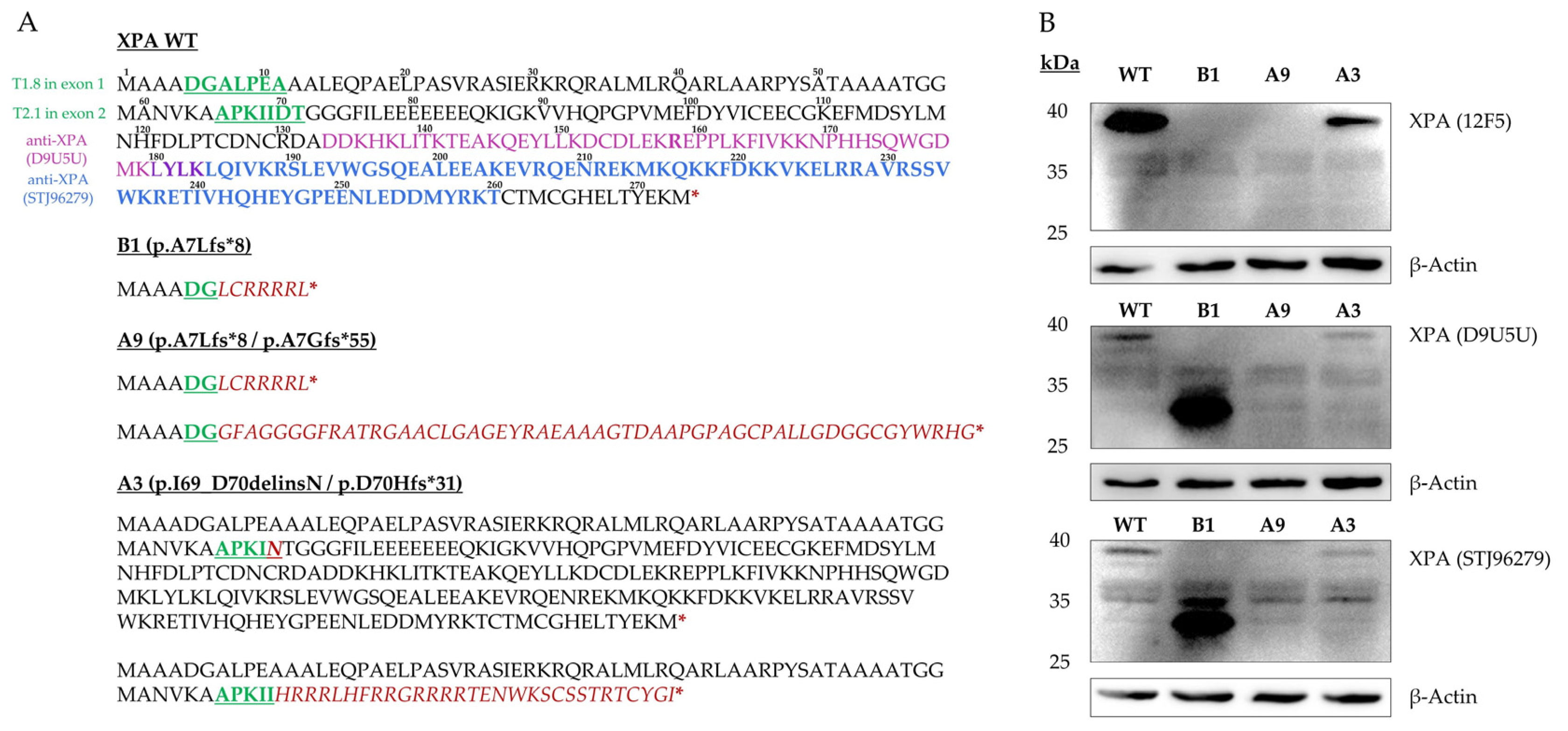
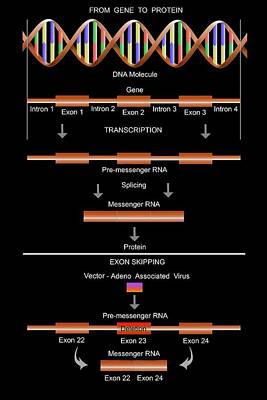
IJMS | Free Full-Text | Homozygous CRISPR/Cas9 Knockout Generated a Novel Functionally Active Exon 1 Skipping XPA Variant in Melanoma Cells | HTML

HnRNP K mislocalisation is a novel protein pathology of frontotemporal lobar degeneration and ageing and leads to cryptic splicing | SpringerLink

SR protein-mediated inhibition of CFTR exon 9 inclusion: molecular characterization of the intronic splicing silencer. - Abstract - Europe PMC

Alternative splicing of α-TM. Exons 2 and 3 are mutually exclusive due... | Download Scientific Diagram

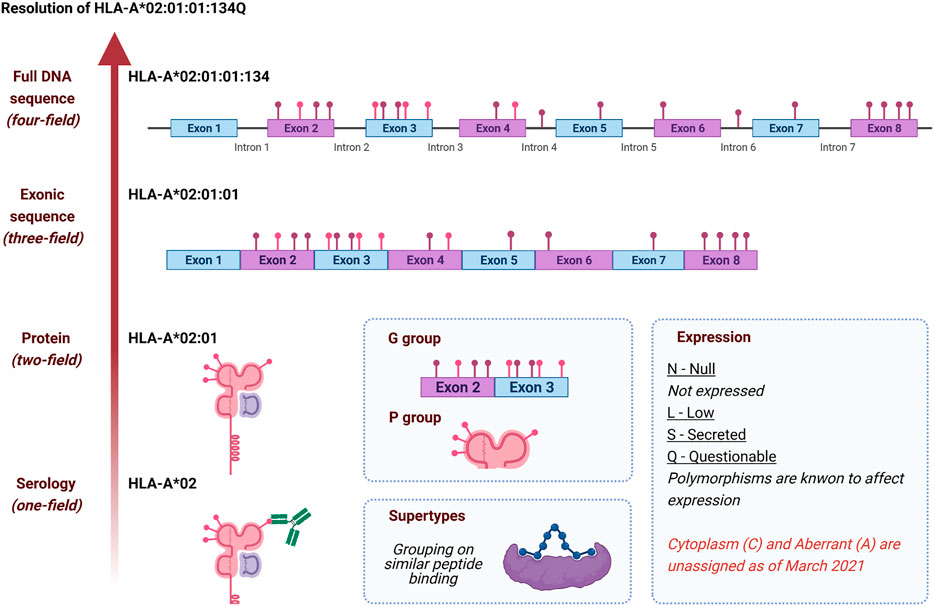
Improved HLA typing of Class I and Class II alleles from next‐generation sequencing data - Sverchkova - 2019 - HLA - Wiley Online Library

Alternative splicing of α-TM. Exons 2 and 3 are mutually exclusive due... | Download Scientific Diagram
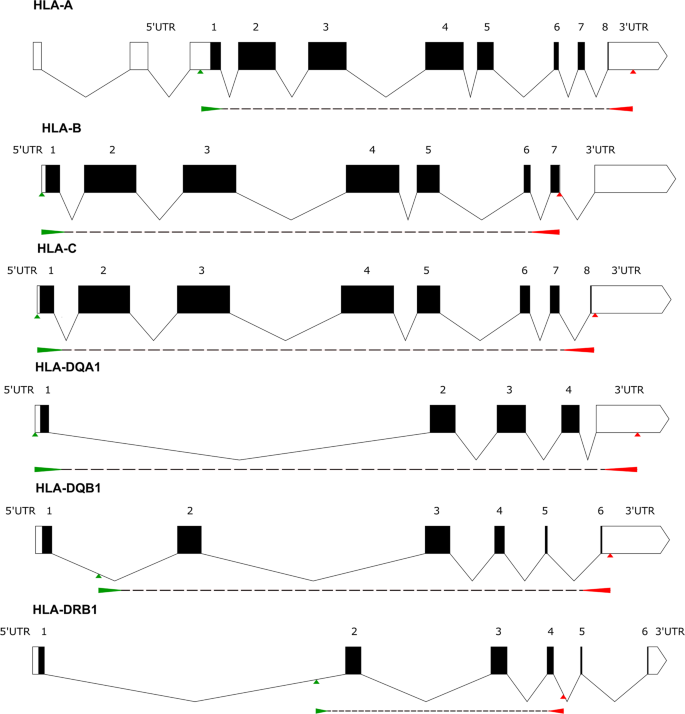
HLA-A, -B, -C, -DRB1, -DQA1, and -DQB1 allele and haplotype frequencies defined by next generation sequencing in a population of East Croatia blood donors | Scientific Reports

Project: Population genetics, anthropology and evolution > 18th International HLA & Immunogenetics Workshop

PCB126 exposure revealed alterations in m6A RNA modifications in transcripts associated with AHR activation | bioRxiv
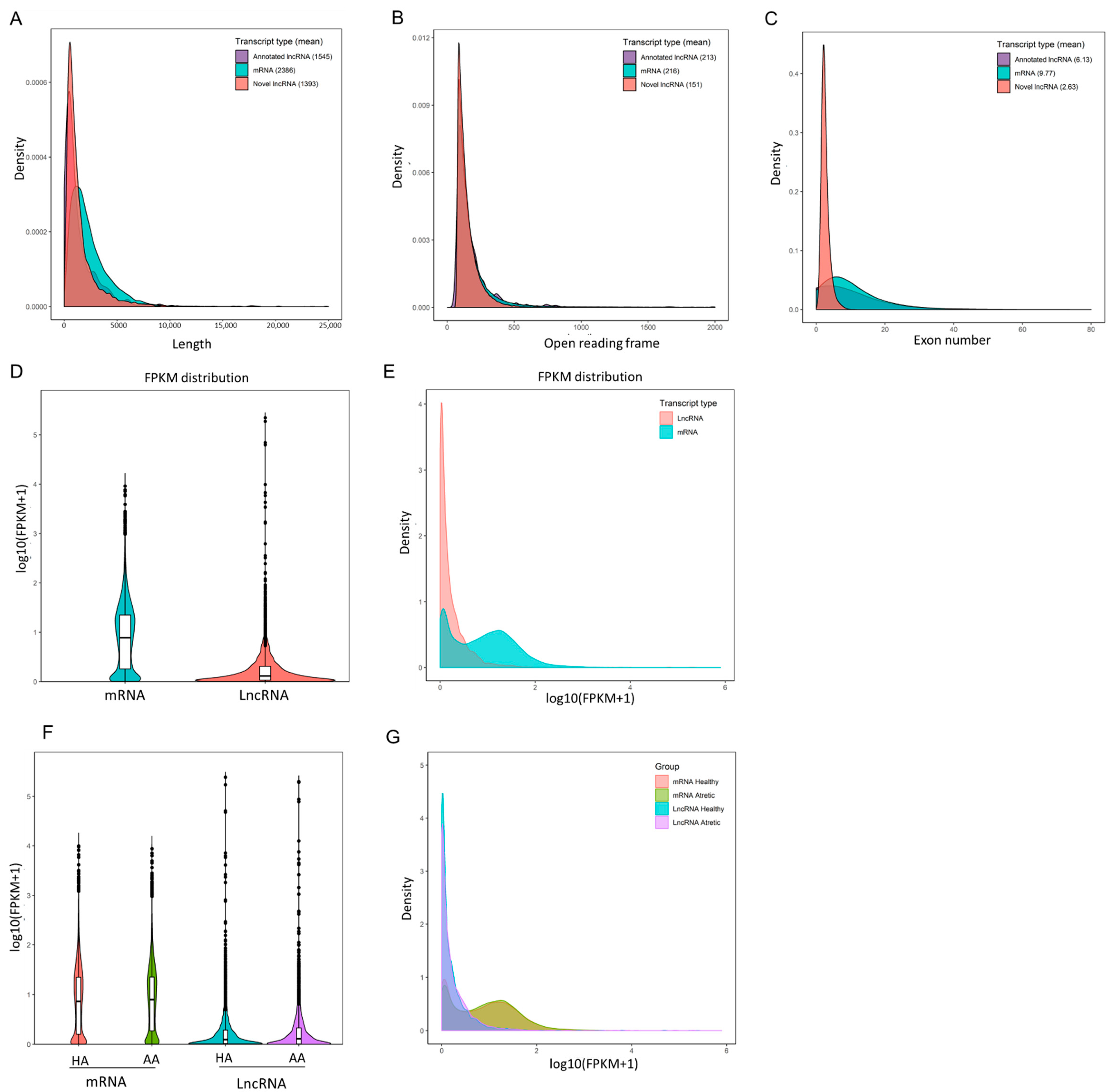
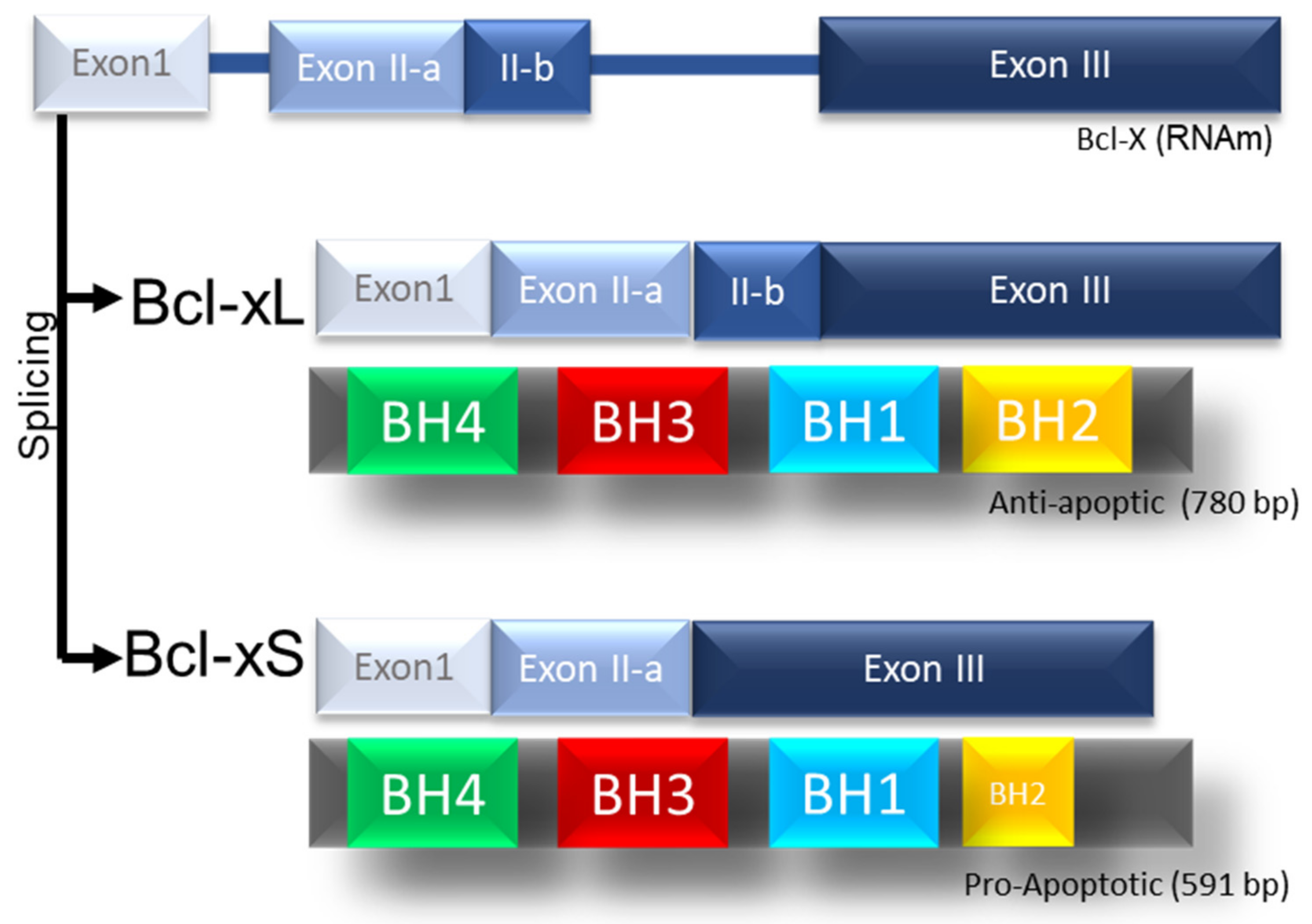
IJMS | Free Full-Text | Characterization of Long Non-Coding RNA Profiles in Porcine Granulosa Cells of Healthy and Atretic Antral Follicles: Implications for a Potential Role in Apoptosis | HTML

Mouse models of neurodegeneration: Know your question, know your mouse | Science Translational Medicine